Project Details:
At the National Cancer Institute, we have demonstrated that higher levels of moderate to vigorous intensity physical activity are associated with a lower risk of cancer, including cancer in the breast, colon, endometrium, bladder, kidney, and stomach1. However, due to a reliance on self-reported measures of physical activity, a number of key questions remain unanswered on what overall volume of physical activity, and what types of physical activity, are associated with lower cancer risk. In addition, previous studies are observational by nature and are therefore unable to determine causality due to unmeasured or residual confounding.
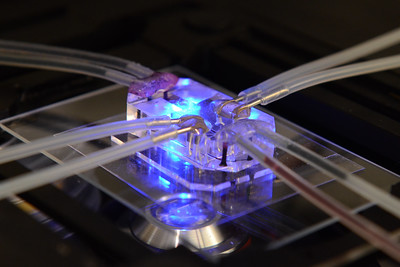
At Oxford, our group has shown that wearable sensors such as wrist-worn accelerometers can be used to noninvasively measure physical activity status in large-scale biomedical studies. For example, we have measured physical activity status in 103,712 UK Biobank participants who agreed to wear a wrist-worn accelerometer for seven days2. These measurements are now actively used by health researchers worldwide to demonstrate that simple measures of overall activity are cross-sectionally associated with cancer outcomes3. However, no large study of device measured physical activity has yet taken place to assess associations with incident cancer outcomes with sufficient longitudinal follow-up. Furthermore, activity trackers often capture ~180 million data points/participant/week and therefore have the potential to identify other powerful behavioural signals to detect future cancer risk.
Machine learning methods can help maximise the utility of data from wearable sensors. These methods attempt to automatically detect patterns in data and then use those uncovered patterns to predict future data. Our group has demonstrated the utility of supervised machine learning to identify sleep and functional physical activity behaviours from raw accelerometer data4. However, there is a broad concern around the lack of reproducibility of machine learning models in health data science5. It is therefore important to carefully consider how to promote robust machine learning findings and reject irreproducible ones, to ensure credibility and trustworthiness.
This DPhil project therefore proposes to use the world’s largest available datasets to investigate what types of physical activity are associated with a lower incidence of cancer. Working with colleagues at the University of Oxford and the National Cancer Institute, you will have the opportunity to address the following important questions:
1. What behavioural measurements of physical activity status can be reliably ascertained from accelerometer datasets?
You will have the opportunity to develop reproducible machine learning skills to develop methods to identify physical activity behaviours from raw accelerometer datasets. Specifically, you will develop semi-supervised machine learning methods which seek to combine supervised methods (good quality labels, small datasets) with unsupervised methods (no labels but large datasets which are less prone to sampling bias). This will involve use of the largest available accelerometer datasets with reference measurements for physical activity behaviours in free-living environments (using wearable cameras)6.
2. What physical activity behaviours are associated with incident cancer events?
Here, you will have the opportunity to develop new skills in epidemiological data analysis. You will have the opportunity to use the UK Biobank dataset which has collected wrist worn accelerometer data from 103,712 participants2. This dataset includes information on participants’ first hospital admission or death from cancer, identified from linkages to the national death index, Hospital Episode Statistics, and cancer registries.
3. Are physical activity behaviours potentially causally associated with cancer?
You will have the opportunity to develop genetic epidemiology skills by implementing two-sample Mendelian Randomization7 to assess potential causal effects of accelerometer measured physical activity and cancer. For cancer outcomes, summary genetic association data will be obtained from existing collaborators from International cancer consortia.
Candidates should have a BSc, or ideally MSc, in a discipline with a substantive epidemiological, computational, or quantitative component. We very much welcome prospective candidates to directly contact us to further develop this proposal.